[1]:
%matplotlib inline
OOB Errors for Random Forests¶
The RandomForestClassifier is trained using bootstrap aggregation, where each new tree is fit from a bootstrap sample of the training observations \(z_i = (x_i, y_i)\). The out-of-bag (OOB) error is the average error for each \(z_i\) calculated using predictions from the trees that do not contain \(z_i\) in their respective bootstrap sample. This allows the RandomForestClassifier to be fit and validated whilst being trained [1].
The example below demonstrates how the OOB error can be measured at the addition of each new tree during training. The resulting plot allows a practitioner to approximate a suitable value of n_estimators at which the error stabilizes.
| [1] |
|
Note: this notebook is a modified version of https://scikit-learn.org/stable/auto_examples/ensemble/plot_ensemble_oob.html to add Randomer Forest and run a little faster.
[2]:
from timeit import default_timer as timer
import matplotlib.pyplot as plt
plt.rcParams['figure.dpi'] = 200
from collections import OrderedDict
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
from rerf.rerfClassifier import rerfClassifier
# Author: Kian Ho <hui.kian.ho@gmail.com>
# Gilles Louppe <g.louppe@gmail.com>
# Andreas Mueller <amueller@ais.uni-bonn.de>
#
# License: BSD 3 Clause
print(__doc__)
RANDOM_STATE = 123
# Generate a binary classification dataset.
X, y = make_classification(n_samples=500, n_features=25,
n_clusters_per_class=1, n_informative=15,
random_state=RANDOM_STATE)
# NOTE: Setting the `warm_start` construction parameter to `True` disables
# support for parallelized ensembles but is necessary for tracking the OOB
# error trajectory during training.
ensemble_clfs = [
("sklearn (Warm start)",
RandomForestClassifier(n_estimators=100,
warm_start=True, oob_score=True,
max_features="sqrt",
random_state=RANDOM_STATE)),
("sklearn (Cold start)",
RandomForestClassifier(n_estimators=100,
warm_start=False, oob_score=True,
max_features="sqrt",
random_state=RANDOM_STATE)),
("ndRF (Base)",
rerfClassifier(n_estimators=100, max_features="sqrt", projection_matrix="Base",
oob_score=True,
random_state=RANDOM_STATE)),
("ndRerF (RerF)",
rerfClassifier(n_estimators=100, max_features="sqrt",
oob_score=True,
random_state=RANDOM_STATE)),
]
# Map a classifier name to a list of (<n_estimators>, <error rate>) pairs.
error_rate = OrderedDict((label, []) for label, _ in ensemble_clfs)
run_time = OrderedDict((label, []) for label, _ in ensemble_clfs)
# Range of `n_estimators` values to explore.
min_estimators = 15
max_estimators = 200
for label, clf in ensemble_clfs:
for i in range(min_estimators, max_estimators + 1, 5):
clf.set_params(n_estimators=i)
start_time = timer()
clf.fit(X, y)
run_time[label].append((i,timer() - start_time))
# Record the OOB error for each `n_estimators=i` setting.
oob_error = 1 - clf.oob_score_
error_rate[label].append((i, oob_error))
Automatically created module for IPython interactive environment
[3]:
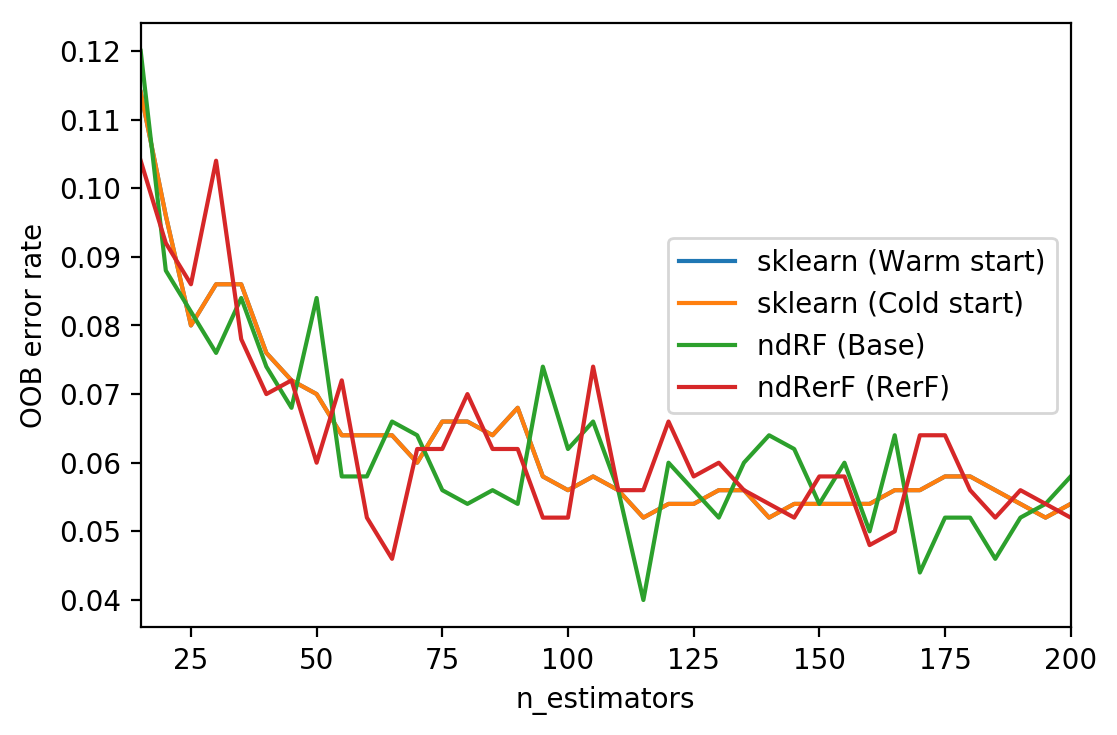
# Generate the "OOB error rate" vs. "n_estimators" plot.
for label, clf_err in error_rate.items():
xs, ys = zip(*clf_err)
plt.plot(xs, ys, label=label)
plt.xlim(min_estimators, max_estimators)
plt.xlabel("n_estimators")
plt.ylabel("OOB error rate")
plt.legend(loc="right")
plt.show()
[4]:
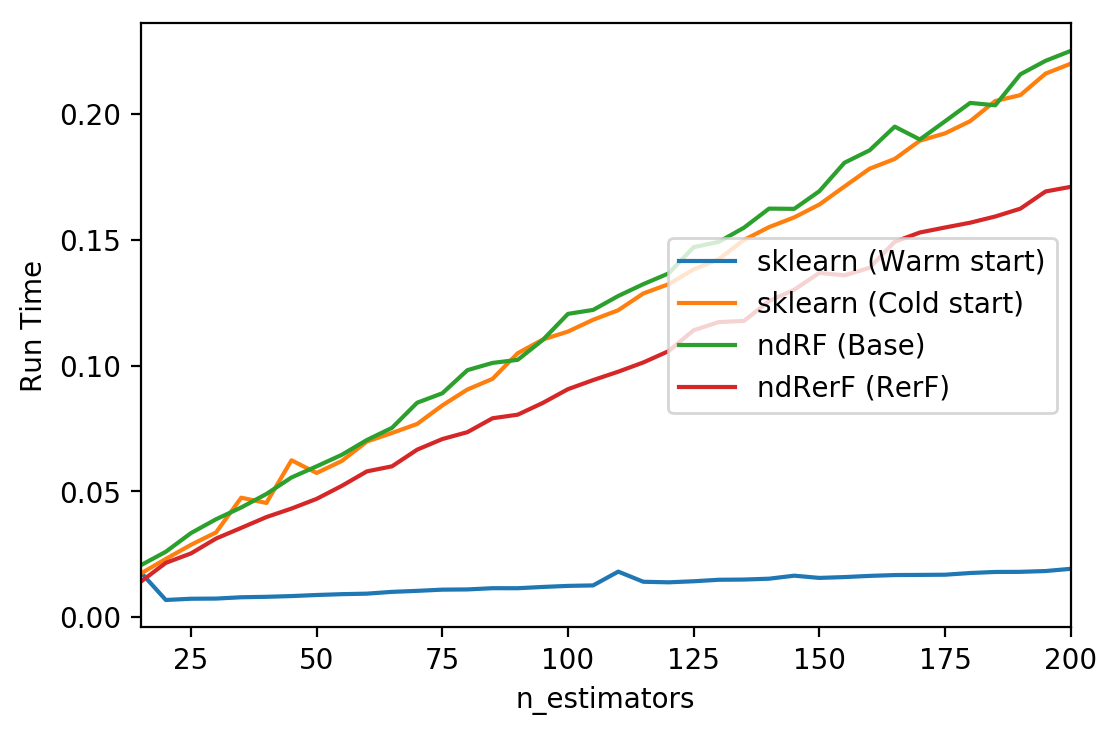
# generate test time plots
for label, rt in run_time.items():
xs, ys = zip(*rt)
plt.plot(xs, ys, label=label)
plt.xlim(min_estimators, max_estimators)
plt.xlabel("n_estimators")
plt.ylabel("Run Time")
plt.legend(loc="right")
plt.show()